the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Spatial gradients in the characteristics of soil-carbon fractions are associated with abiotic features but not microbial communities
Julia Indivero
Cailene Gunn
Malak M. Tfaily
Rosalie K. Chu
Jason Toyoda
Vanessa L. Bailey
Nicholas D. Ward
James C. Stegen
Coastal terrestrial–aquatic interfaces (TAIs) are dynamic zones of biogeochemical cycling influenced by salinity gradients. However, there is significant heterogeneity in salinity influences on TAI soil biogeochemical function. This heterogeneity is perhaps related to unrecognized mechanisms associated with carbon (C) chemistry and microbial communities. To investigate this potential, we evaluated hypotheses associated with salinity-associated shifts in organic C thermodynamics; biochemical transformations; and nitrogen-, phosphorus-, and sulfur-containing heteroatom organic compounds in a first-order coastal watershed on the Olympic Peninsula of Washington, USA. In contrast to our hypotheses, thermodynamic favorability of water-soluble organic compounds in shallow soils decreased with increasing salinity (43–867 µS cm−1), as did the number of inferred biochemical transformations and total heteroatom content. These patterns indicate lower microbial activity at higher salinity that is potentially constrained by accumulation of less-favorable organic C. Furthermore, organic compounds appeared to be primarily marine- or algae-derived in forested floodplain soils with more lipid-like and protein-like compounds, relative to upland soils that had more lignin-, tannin-, and carbohydrate-like compounds. Based on a recent simulation-based study, we further hypothesized a relationship between C chemistry and the ecological assembly processes governing microbial community composition. Null modeling revealed that differences in microbial community composition – assayed using 16S rRNA gene sequencing – were primarily the result of limited exchange of organisms among communities (i.e., dispersal limitation). This results in unstructured demographic events that cause community composition to diverge stochastically, as opposed to divergence in community composition being due to deterministic selection-based processes associated with differences in environmental conditions. The strong influence of stochastic processes was further reflected in there being no statistical relationship between community assembly processes (e.g., the relative influence of stochastic assembly processes) and C chemistry (e.g., heteroatom content). This suggests that microbial community composition does not have a mechanistic or causal linkage to C chemistry. The salinity-associated gradient in C chemistry was, therefore, likely influenced by a combination of spatially structured inputs and salinity-associated metabolic responses of microbial communities that were independent of community composition. We propose that impacts of salinity on coastal soil biogeochemistry need to be understood in the context of C chemistry, hydrologic or depositional dynamics, and microbial physiology, while microbial composition may have less influence.
- Article
(4297 KB) - Full-text XML
-
Supplement
(1087 KB) - BibTeX
- EndNote
The interface between terrestrial and aquatic ecosystems represents a dynamic and poorly understood component of the global carbon (C) cycle, particularly along the tidally influenced reaches of coastal watersheds where terrestrial and marine biospheres intersect (Krauss et al., 2018; Neubauer et al., 2013; Tank et al., 2018; N. D. Ward et al., 2017). Moreover, the nutrient cycles occurring at these terrestrial–aquatic interfaces (TAIs) influence locally important ecosystem services like contaminant fate and transport and water quality (Conrads and Darby, 2017; Vidon et al., 2010). While coastal soil-C stocks are being increasingly quantified (Hinson et al., 2017; Holmquist et al., 2018; Krauss et al., 2018), the impact of tidally driven salinity gradients on molecular-level features of the soil-C pool and the processes driving soil organic matter (OM) cycling are poorly studied (Barry et al., 2018; Hoitink et al., 2009; Sawakuchi et al., 2017; N. D. Ward et al., 2017). This is particularly true in settings with low freshwater inputs that allow for significant seawater intrusion compared to large river systems (Hoitink and Jay, 2016).
While multiple processes impact TAI carbon pools (e.g., tidal inputs, in situ root exudates and litter inputs, and decomposition processes), there is some indication that microbial diversity and composition impact soil-C storage and mineralization (Mau et al., 2015; Trivedi et al., 2016). This points to the intriguing possibility that processes governing microbial community assembly may be associated with OM chemistry, but evaluations of such associations are lacking. This lack of mechanistic knowledge combined with significant ecosystem heterogeneity in biogeochemical function across salinity gradients (more below) highlights a need to understand how molecular-level processes vary with seawater exposure along coastal TAIs. Doing so will help enhance predictive models of TAI biogeochemistry that can be potentially included in ecosystem models to more accurately represent the role of TAIs in the broader Earth system (U.S. DOE, 2017).
Modeling of coastal TAIs is currently impeded by poor knowledge of the mechanisms underlying salinity-driven variation in biogeochemical function of associated soils. Previous studies have evaluated function primarily as carbon dioxide (CO2) and methane (CH4) flux measurements from soil and/or soil OM concentrations measured as bulk soil C, percent OM, and porewater dissolved organic C (DOC) concentrations in large-scale coastal plain river systems. Results from field-based natural salinity gradient studies, long-term field manipulations of salinity exposure, and lab-based incubation studies subjecting soils to varying levels of salinity broadly show the following trends: increases in CO2 and decreases in CH4 emissions in freshwater soils exposed to increasing salinity (Chambers et al., 2011, 2013, 2014; Liu et al., 2017; Marton et al., 2012; Neubauer et al., 2013; Steinmuller and Chambers, 2018; Weston et al., 2006, 2011) and negative relationships between salinity and emissions of both CO2 and CH4 from soils with a history of exposure to high salinity (Chambers et al., 2013; Herbert et al., 2018; Neubauer et al., 2005, 2013; Weston et al., 2014; also see Table S1 in the Supplement). Exceptions have been observed where CO2 emissions decreased in historically freshwater coastal wetland soils exposed to seawater (Ardón et al., 2018; Herbert et al., 2018), with Ardón et al. (2018) also reporting an increase in CH4 flux with salinity and Steinmuller and Chambers (2018) reporting no change in CH4 flux with increasing salinity. These observations suggest that microbial activity usually increases with salinity in soils that were not previously exposed to saline conditions while simultaneously indicating reduced microbial activity with increasing salinity in soils that have a historical exposure to high salinity. In contrast to relatively consistent responses of gas fluxes to changes in salinity, there are strong inconsistencies in DOC responses, including no change (Weston et al., 2006, 2011, 2014), increased DOC (Chambers et al., 2014; Tzortziou et al., 2011), and decreased DOC (Ardón et al., 2016, 2018; Liu et al., 2017; Yang et al., 2018) with increasing salinity.
Relatively consistent gas flux responses to changes in salinity combined with inconsistent DOC responses to elevated salinity suggest decoupling between biogeochemical rates and the concentration of DOC. This apparent decoupling between the size of the C pool (in this case the concentration of DOC) and microbial activity suggests that C biogeochemistry is influenced by salinity-exposure history, which in turn influences nutrient resources available to soil microbial communities. Specifically, any systematic shifts in soil organic carbon (SOC) chemistry (along salinity gradients) that cannot be observed with bulk C measurements (e.g., changes in chemistry that reduce C bioavailability; Neubauer et al., 2013) may result in unpredictable carbon fluxes. Moreover, bulk C content can show no change across gradients of salinity (Neubauer et al., 2013) and may fail to capture an integrated view of microbially driven C cycling dynamics at TAIs. In contrast, detailed molecular-level evaluation of SOC composition can provide a more mechanistic view of organic carbon (OC) transformations relative to bulk measures of C content or gas flux measurements.
Despite its potential importance, a detailed understanding of the characteristics of soil organic compounds (Zark and Dittmar, 2018) and their association with microbial communities in coastal TAIs is currently not available. Nonetheless, we derived a series of expectations by first recognizing that (1) our study system is a historically freshwater system, only recently having been exposed to saltwater due to removal of a culvert in 2014 (see Methods), and that (2) microbial activity increases with increasing salinity in historically freshwater systems (Nyman and Delaune, 1991; Smith et al., 1983; Tzortziou et al., 2011). In addition, it is generally expected that microbes preferentially degrade compounds with higher nominal oxidation states (NOSCs) or lower Gibbs free energy () due to greater thermodynamic favorability (Boye et al., 2017; Graham et al., 2017a, 2018; Stegen et al., 2018b; C. P. Ward et al., 2017). An important caveat is that factors such as redox state, physical protection, mineral associations, and microbial community composition can alter this pure chemistry-based expectation (Schmidt et al., 2011). As a simple point of departure, however, we assume that OM reactivity follows NOSCs, thereby leading to our first expectation or hypothesis: the average of OM will increase with increasing salinity as organic compounds with greater thermodynamic favorability are preferentially depleted (LaRowe and Van Cappellen, 2011) due to microbial activity increasing with salinity. To develop our second hypothesis we note that actively growing microbial communities are known to enhance biochemical transformations and generate heteroatom-containing organic molecules (sulfur (S), nitrogen (N) and phosphorus (P)) (Guillemette et al., 2018; Koch et al., 2014; Ksionzek et al., 2016); therefore greater heteroatom content and more biochemical transformations are expected with increasing salinity. Our third hypothesis is based on microorganisms adapting to saline conditions through the production or sequestration of osmolytes (Gouffi et al., 1999; Sleator and Hill, 2002), a strategy that may require microbes to break down organic molecules to extract N (i.e., N mining). We therefore hypothesize increases in N-containing biochemical transformation with increasing salinity. Our fourth hypothesis is based on the observation that soils in saturated environments like floodplains are expected to be less oxygenated and can also receive deposition of marine- or algae-derived OM and suspended sediments during tidal flooding. These factors can result in OM having lower oxygen-to-carbon (O∕C) and higher hydrogen-to-carbon (H∕C) ratios as compared to upland soils (Seidel et al., 2016; Tfaily et al., 2014; Ward et al., 2019b). We therefore hypothesize a greater relative abundance of lipid- and protein-like and less lignin- and tannin-like compounds in the floodplain soils, relative to upland (i.e., drained) soil.
While we expect systematic shifts in C chemistry across landscape-scale salinity gradients, an open question is the degree to which C chemistry is associated with ecological assembly processes governing composition of microbial communities. Soil microorganisms transform soil C, but there is limited evidence of direct links between microbial community assembly processes and molecular-level soil-C chemistry (Kubartová et al., 2015; Rocca et al., 2015; Trivedi et al., 2016; van der Wal et al., 2015). Assembly processes, broadly divided into deterministic (systematic impacts of selection) and stochastic (unstructured demographic events) factors, function over space and time to influence the composition of microbial communities, which in turn mediate biogeochemical cycles (Graham et al., 2016, 2017b; Nemergut et al., 2013; Stegen et al., 2015). Deterministic processes lead to selection of microbial communities resulting from different organisms having different levels of fitness for a given set of environmental conditions, including abiotic variables and biotic factors related to organismal interactions, while stochastic processes include random birth or death events and unstructured dispersal. The relative influences of stochastic and deterministic processes can be inferred from phylogenetic distances among microbial taxa using ecological null models. This approach has been widely employed to understand community assembly processes in surface and subsurface microbial ecology (Caruso et al., 2011; Dini-Andreote et al., 2015; Graham et al., 2017a, 2018; Sengupta et al., 2019; Stegen et al., 2012). Furthermore, a recent study used ecological simulation modeling to show that communities experiencing increased rates of dispersal are linked to reduced biogeochemical functioning (Graham and Stegen, 2017). This, combined with evidence of increasing microbial activity with increasing salinity (discussed above), leads to a fifth hypothesis that the influence of deterministic selection will progressively increase with salinity due to increased microbial activity.
Analyses of specific chemical biomarkers such as lignin phenols, amino acids, and lipids have been used in soils, sediments, and water to quantitatively evaluate the provenance of terrestrial-derived OM (Hedges et al., 1997), the reactivity of OM as it travels through a soil column (Shen et al., 2015), and microbial community composition (Langer and Rinklebe, 2009), respectively. While biomarkers provide quantitative details on OC cycling, they generally represent a small fraction of the total OM pool; thus, non-targeted approaches such as analysis of thousands of organic molecules via Fourier-transform ion-cyclotron-resonance mass spectrometry (FTICR-MS) have become increasingly widespread for determining molecular-level organic compound signatures (Rivas-Ubach et al., 2018) across a variety of terrestrial (Bailey et al., 2017; Simon et al., 2018), aquatic or marine (Lechtenfeld et al., 2015), and transitional settings such as hyporheic zones (Graham et al., 2017a; Stegen et al., 2016, 2018b) and river–ocean gradients (Medeiros et al., 2015).
The objective of the current study was to test the following hypotheses in a coastal forested floodplain and adjacent upland forest: (1) mean Gibbs free energy of organic compounds will increase with increasing salinity; (2 and 3) biochemical transformations, heteroatom content, and N-containing biochemical transformation will increase with increasing salinity; (4) lipid- and protein-like compound classes will be more prevalent in the floodplain soils compared to upland soils in which lignin- and tannin-like molecules will dominate; and (5) microbial community assembly processes will be increasingly deterministic as salinity increases. The chemical forms of C in these soils were characterized using FTICR-MS. We also employed ecological null-model analysis to evaluate the relationship between C chemistry and the influences of assembly processes on microbial communities. Based on our results, we propose a conceptual model of organic C processing in a coastal forested floodplain in which landscape-scale gradients in C chemistry are driven by a combination of spatially structured inputs and salinity-associated metabolic responses of microbial communities that are independent of community composition.
2.1 Site information and soil sampling
Soils along a coastal watershed draining a small first-order stream, Beaver Creek, on the Washington coast were selected for this study (lat 46.907, long −123.976). Beaver Creek is a tributary of Johns River and experiences a high tidal range of up to 2.5 m that extends midway up the first-order stream's channel and inundates the landscape in its floodplains. The confluence of Beaver Creek and Johns River is roughly 2.5 km upstream of the Grays Harbor estuary and 14.5 km from the Pacific Ocean, and it experiences variable exposure to saline waters at high tide (Fig. 1). Surface water salinity near Beaver Creek's confluence ranges from 0 PSU (PSU – practical salinity unit) at low tide to 30 PSU at high tide during dry periods (Nicholas D. Ward, unpublished data). The Beaver Creek watershed is 3.8 km2. The tidal floodplain makes up 0.5 km2 of this total watershed area. Tidal exchange to Beaver Creek was restored after 2014, when a culvert near the creek's confluence with Johns River was removed (Washington Department of Fish and Wildlife, 2019). Due to the minimal past tidal exchange, the floodplain is dominated by gymnosperm trees (Picea sitchensis) that have been rapidly dying since the culvert removal (Ward et al., 2019a). The headwaters (before the river channel forms) are a sparsely forested, perennially inundated freshwater wetland with tidal exchange blocked by a beaver dam, followed downstream by a densely forested setting along the river channel. Towards Beaver Creek's confluence, salt-tolerant grasses such as Agrostis stolonifera (creeping bentgrass) become the most dominant land cover as forest cover becomes more sparse. The watershed hillslope and uplands are dominated by Tsuga heterophylla (western hemlock) trees, but Picea sitchensis (Sitka spruce) are also present.
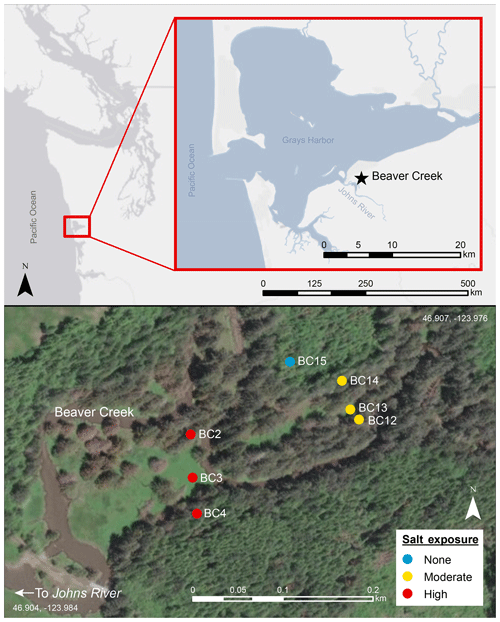
Figure 1Study site Beaver Creek on the Olympic Peninsula in western Washington. The creek is a first-order stream, with tidal exchange restored in 2014. Top panel shows site location in western Washington, with inset panel zoomed in to show the site close to Johns River. Bottom panel shows soil sampling locations at the high salt-exposure (BC2, BC3, and BC4) transect, moderate salt-exposure (BC12, BC13, and BC14) transect, and terrestrial upland (BC15) site. The transects with six sampling sites experience periodic inundation episodes which result in surface pooling of tidal water. Map was created using ArcGIS 10.5 software (ESRI, 2017). Coordinate system: GCS WGS 1984. The service layer is credited to Esri, HERE, Garmin©, OpenStreetMap© contributors, and the GIS user community, with resources distributed under a Creative Commons BY-SA License.
Two sampling transects perpendicular to the river along the upstream–downstream salinity gradient were established and represent a high salt exposure site close to the culvert breach location and a moderate salt-exposure site upstream of the high salt exposure site. These transects represent a coastal forested wetland with brackish (semi-salty) groundwater and consisted of three terrestrial sampling points at each transect extending from the riparian zone to the beginning of the steep upslope. An additional soil sampling point ∼20 m uphill from the moderate salt-exposure site transect served as a purely terrestrial upland endmember. The soils are Andisols, and floodplain transects represented hydric soils classified as Ocosta silty clay loam, while the upland site was a well-drained Mopang silt loam. The transects experience periodic inundation episodes which result in surface pooling of tidal water, which can be up to ∼1 m deep. The water table varies seasonally and during tidal cycles and inundation events, ranging from about 0 to 1 m below the ground surface (Nicholas D. Ward, unpublished data).
Soil samples were collected in triplicate at each of the seven locations (Fig. 1; BC2 (2.94 m), BC3 (2.63 m), and BC4 (2.82 m) at the high salt-exposure transect; locations BC12 (2.82 m) , BC13 (2.67 m), and BC14 (3.07 m) at the moderate salt-exposure transect; and BC15 (13.45 m) as the upland site). The high salt-exposure transect was 230 m from the moderately saline transect (0.6 km from the confluence of Beaver Creek with Johns River), and each site at the transect was ∼35 m apart from the next. Each transect is ∼90 m. For data comparison's sake, we classify BC2, BC3, BC12, and BC13 as floodplain sites, while BC4 and BC14 are further inland and ∼75 m away from the creek at the base of the densely wooded hillslope. Soil samples for molecular characterization studies were collected at two depths – shallow (10 cm) and deep (19–30 cm). Samples were collected from the face of soil pits using custom mini-corers, placed into sterile amber glass vials, purged with N2 to maintain anaerobic conditions, frozen in the field within an hour at −20 ∘C, and stored at −80 ∘C on return to the lab. Bulk samples were collected for soil physicochemical characterization, including texture classification with the hydrometer method after organic matter removal; dry combustion with direct measure of total C, nitrogen (N), and sulfur (S) by the Elementar MACRO Cube; plant-available N as ammoniacal nitrogen (NH4-N) and nitrate nitrogen (NO3-N) with 2 M KCl quantified on Lachat QuikChem 8500 Series 2 (Hach, Loveland, Colorado) as a colorimetric reaction; pH measured as 1:2 soil : water slurry measured with a Hanna benchtop meter; specific conductivity measured as 1:5 soil : water slurry measured on an MP-6p portable specific conductivity meter; gravimetric water content measured after drying soils at 105 ∘C for 48 h; and bulk density and porosity measured as per standard methods (Sumner, 1999). Soil chemical characterization was performed on air-dried sieved soils.
2.2 FTICR-MS solvent extraction and data acquisition
Soil organic compounds were extracted using a sequential extraction protocol with polar {water (H2O)} and non-polar {chloroform (CHCl3)(representing mineral-bound fraction)} solvents per standardized protocols (Graham et al., 2017a; Tfaily et al., 2015, 2017), which extract about 2 %–15 % of total organic carbon and represent both polar and non-polar soil organic carbon fractions. Importantly, our analyses do not depend on extracting a large portion of the C found within a given soil sample. Instead, we assume that the extracted fraction is a representative sub-sample. This is a standard approach and assumption made in any study examining metabolites or other types of organic molecules in soil. Briefly, extracts were prepared by adding 5 mL of Milli-Q H2O to 5 g of each of the replicate samples in sterile polypropylene centrifuge tubes (Genesee Scientific, San Diego, USA) suitable for organic solvent extractions and shaking for 2 h on a Thermo Scientific LP Vortex Mixer. Samples were removed from the shaker and centrifuged for 5 min at 6000 rpm, and the supernatant was removed into a fresh centrifuge tube. This step was repeated two more times, with the 15 mL supernatant pooled for each sample and stored at −80 ∘C until further processing. Next, Folch extraction with CHCl3 and CH3OH was performed for each soil pellet left over from the water extraction. Folch extraction entailed adding 2 mL CH3OH, vortexing for 5 s, adding 4 mL of CHCl3, vortexing for 5 s, and adding 0.25 mL of Milli-Q H2O. The samples were shaken for 1 h, and another 1.25 mL of Milli-Q H2O was added and left overnight at 4 ∘C to obtain bi-layer separation of the upper (polar) layer and the lower (non-polar) layer. The extracts were stored in glass vials at −20 ∘C until they were ready to be used. The water-soluble organic carbon (WSOC) fraction was further purified using a sequential phase extraction (SPE) protocol to remove salts as per Dittmar et al. (2008). Briefly, samples were acidified to a pH of 2 with 85 % phosphoric acid. The samples were passed through Bond Elut PPL cartridges (©Agilent Technologies) that were preactivated with CH3OH. The cartridges were washed five times with 10 mM of HCl, which was followed by nitrogen-gas drying. Next, 1.5 mL of CH3OH, a solvent that is compatible with direct analysis on the FTICR-MS, was used to elute the samples from the cartridge, thus avoiding an additional evaporation step and reducing the chance of losing volatile organic compounds and saving time during sample preparation. While SPE by PPL has been shown not to be very effective in extracting several major classes of DOM compounds that had high ESI efficiencies, such as carboxylic acids and organo-sulfur compounds, and that out-competed other less-functionalized compounds (e.g., carbohydrates) for charge in the ESI source (Tfaily et al., 2012), it is highly efficient for marine and estuary DOM samples, as it provides complete desalination of the sample. Loss of small molecules such as simple sugars is known to happen during SPE; however this is not a concern for the current study, as FTICR-MS is sensitive to compounds above 200 Da. In this study, SPE by PPL isolates a major DOM fraction that is salt-free, allowing for DOM characterization by FTICR-MS (Dittmar et al., 2008). While we did not measure SPE extraction efficiency for this study, it usually ranges between 40 % and 62 % depending on the sample (Dittmar et al., 2008). Samples that are collected from the same ecosystem have been shown to have similar extraction efficiency. The extracts were injected into a 12 Tesla Bruker SolariX FTICR-MS located at Environmental Molecular Sciences Laboratory (EMSL) in Richland, Washington, USA. Detailed methods for instrument calibration, experimental conditions, and data acquisition are provided in Graham et al. (2017a) and Tfaily et al. (2017).
2.3 FTICR-MS data processing
One hundred forty-four individual scans were averaged for each sample and internally calibrated using an organic matter homologous series separated by 14 Da (–CH2 groups). The mass measurement accuracy was less than 1 ppm for singly charged ions across a broad m∕z range (m∕z 100–900). Data analysis software (Bruker Daltonik version 4.2) was used to convert raw spectra to a list of m∕z values, applying the FTMS peak picker module with a signal-to-noise ratio (S∕N) threshold set to 7 and absolute intensity threshold set to the default value of 100. Chemical formulae were then assigned using in-house software following the compound identification algorithm that was proposed by Kujawinski and Behn (2006), modified by Minor et al. (2012), and described in Tolić et al. (2017). Peaks below 200 and above 900 were dropped to select only for calibrated and assigned peaks. Chemical formulae were assigned based on the following criteria: and mass measurement error < 0.5 ppm, taking into consideration the presence of C, H, O, N, S, and P and excluding other elements. Detected peaks and the associated molecular formula were uploaded to the in-house pipeline FTICR R Exploratory Data Analysis (FREDA) to obtain (i) NOSC values from elemental composition of the organic compounds (Koch and Dittmar, 2006, 2016), (ii) thermodynamic favorability of the compounds calculated as Gibbs free energy for the oxidation half-reactions of the organic compounds () based on the equation (LaRowe and Van Cappellen, 2011), where a higher indicates a less thermodynamically favorable species than a lower value (LaRowe and Van Cappellen, 2011), (iii) abundance of compounds grouped into elemental groups (CHO, CHOS, CHOP, CHNOS, CHNO, CHNOP, CHOSP, and CHNOSP), and (iv) abundance of compound classes (carbohydrate-, lipid-, protein-, amino-sugar-, lignin-, tannin-, condensed hydrocarbon-, and unsaturated hydrocarbon-like) based on molar H:C and O:C ratios of the compounds (Bailey et al., 2017).
Biochemical transformations potentially occurring in each sample were inferred from the FTICR-MS data by comparing mass differences in peaks within each sample to precise mass differences for commonly observed biochemical transformations (Breitling et al., 2006; Stegen et al., 2018b). The ultra-high mass accuracy of FTICR-MS allows precise mass differences to be counted for the number of times each transformation was observed within each sample. Briefly, the mass difference between m∕z peaks extracted from each spectrum was compared to commonly observed mass differences associated with 92 common biochemical transformations provided in previous publications (Graham et al., 2017a; Stegen et al., 2018b). All possible pairwise mass differences were calculated within each extraction type for each sample, as shown in Fig. 2, where a mass difference of 2.01586 indicates a hydrogenation reaction. It is important to note that the direct-injection electrospray ionization (ESI) FTICR-MS approach cannot distinguish between isomers, such as in the case of a mass difference corresponding to a loss of grain of glucose, fructose, or galactose.
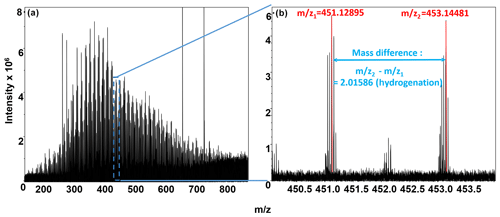
Figure 2(a) Negative-mode FTICR-MS (full spectrum); (b) magnification at m∕z ∼450, showing an example of our FTICR-MS spectra, overlain with peak mass assignments (red), and a biochemical transformation (mass difference between peaks; denoted in blue). y axis denotes peak intensities, and x axis denotes mass-to-charge ratio.
2.4 Ecological modeling
Null modeling was used to estimate influences of ecological processes on microbial community composition from rarefied (10 000) 16S rRNA amplicon data, providing estimates of microbial community composition and phylogenetic relatedness. The extraction, purification, and sequencing of soil microbial DNA were performed according to published protocol (Bottos et al., 2018). Briefly, microbial DNA was extracted from 0.25 g of each sample using the MoBio PowerSoil DNA Isolation Kit and cleaned up using a Zymo ZR-96 Genomic DNA Clean and Concentrator-5 kit (Zymo Research Corporation, Irvine, CA) as per manufacturer instructions. The V4 region of the 16S rRNA gene was amplified by a polymerase chain reaction and amplicons sequenced on Illumina MiSeq using the 500 MiSeq Reagent Kit v2 (Illumina Inc., San Diego, CA) according to the manufacturer's instructions. Sequence pre-processing, operational taxonomic unit (OTU) table construction, and phylogenetic tree building were performed using an in-house pipeline, Hundo (Brown et al., 2018). Briefly, sequence demultiplexing was done using ea-utils (Aronesty, 2013), reads quality filtered with BBDuk2 (Bushnell, 2018), and was merged using USEARCH (Edgar, 2010). Sequence de-replication and clustering was performed, taxonomy was assigned to the operational taxonomic unit (OTU), and chimeras were removed using USEARCH. Raw sequences are archived at NCBI (BioProject PRJNA541992) at the following website: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA541992 (last access: 7 October 2019).
Null modeling was performed as described previously (Stegen et al., 2013, 2015), with a total of 35 samples to estimate relative influences of deterministic and stochastic selection processes. Briefly, samples that passed quality control and a rarefaction threshold were evaluated for pairwise phylogenetic turnover between communities, calculated as the difference between the observed values of the β-mean nearest taxon distance (βMNTD) and mean of the null βMNTD distribution in units of standard deviation (see Stegen et al., 2012, for details). The difference between observed βMNTD and the null distribution is known as the β-nearest taxon index (βNTI). Deterministic assembly processes are assumed to be dominant when βNTI > 2 or . When βNTI is >2 it indicates that deterministic processes have driven community composition apart, which is referred to as “variable selection” (Dini-Andreote et al., 2015; Stegen et al., 2015). When βNTI is it indicates that deterministic processes have caused community composition to be similar between a given pair of communities, which is referred to as “homogeneous selection” (Dini-Andreote et al., 2015; Stegen et al., 2015).
Pairwise community comparisons that do not deviate significantly from the null distribution (i.e., ) indicate the dominance of stochastic processes (including homogenizing dispersal and dispersal limitation) or a scenario in which neither deterministic or stochastic processes dominate (referred to as undominated). Homogenizing dispersal occurs when the rate of dispersal between two communities results in community composition becoming relatively similar between the two communities and potentially overwhelming other assembly processes (e.g., variable selection). Dispersal limitation is the result of very low rates of organismal exchange between communities, which can result in the stochastic divergence of community composition through the accumulated outcomes of random birth or death events (i.e., ecological drift).
For pairwise comparisons that were not associated with deterministic processes (i.e., when ), we use a second null model to test for influences of homogenizing dispersal or dispersal limitation. This second null model is referred to as RCbray and, like βNTI, accounts for variation in OTU relative abundances (Stegen et al., 2013, 2015). Homogenizing dispersal was assumed to be the dominant process for pairwise comparisons characterized by and RC. Dispersal limitation was assumed to be the dominant process for pairwise comparisons characterized by and RCbray>0.95. The relative influences of variable selection, homogeneous selection, dispersal limitation, and homogenizing dispersal were quantified by the fraction of pairwise comparisons that were dominated by each ecological process (Stegen et al., 2013). The relative contribution of scenarios in which the system was undominated was estimated as the fraction of pairwise comparisons characterized by and (Stegen et al., 2015).
2.5 Statistical methods
Samples were separately analyzed for WSOC and CHCl3 fractions. Within each solvent fraction, samples were grouped into shallow or deep depths. FTICR-MS-dependent metrics including , and relative abundance of compound classes, total transformations, N-containing transformations, and organic nitrogen containing compounds, were regressed against specific conductivity. Regressions were considered significant if R2≥0.50 and p≤0.05. The transformation profiles were also regressed with the community assembly processes to determine the relationship between deterministic or stochastic processes and organic compound transformations. Mantel tests were used to evaluate similarity between the βNTI matrix and Sørensen matrix of peak presence or absence. The Sørensen distance matrices of WSOC and CHCl3 peaks were regressed against measured variables (soil physicochemical properties) and community assembly process variables to determine correlations. Finally, a redundancy analysis-based stepwise model building with forward model choice was performed to determine variation in the Hellinger-transformed water-fraction peaks and CHCl3 fraction peaks as explained by explanatory variables (which included measured soil variables, modeled community assembly variables, and categorical variables depth and location). All statistical analyses were performed in the statistical computing language R version 3.5.3 (R Development Core Team, 2019).
3.1 Soil characterization
The percent of total soil C (%C) in the shallow soils (26.3±8.3 %) was higher than the deeper soils (4.0±1.3 %) for the lowland soils (i.e., “floodplain” and “inland” sites), while the upland site had an average %C of 7.4±0.27 % at 10 cm and 2.13±0.06 % at 30 cm (Table S2). No significant relation was observed between %C in the shallow inland and floodplain soils along the salinity gradient. The percent of total soil N (%N) of the shallow soils was higher (1.5±0.40 %) than the deeper soils (0.4±0.08 %) for the lowland soils and co-varied with %C (r2=0.95). The pH values of all soils were acidic (5.64±0.70). The concentrations of NH4-N and NO3-N showed a consistent trend, where NH4-N was 1–2 orders of magnitude higher than NO3-N in all samples. The specific conductivity (used as a measurement of salinity in this study) of the shallow soils ranged from 206 to 866 µS cm−1 (±12) in the lowland soils to 43±5 µS cm−1 in the terrestrial endmember site. The deep soils exhibited specific conductivity ranging from 148 to 524 µS cm−1 (±11) in the lowland soils to 29.2±8 µS cm−1 in the terrestrial endmember site. Texture analysis revealed a broad range of sand (4.1 %–40 %), silt (21.4 %–57.9 %), and clay (28.6 %–64.8 %) fractions.
3.2 Thermodynamics, compound classes, and elemental composition
The calculated WSOC (Table S3) in shallow soils was consistent with our hypothesis of decreasing thermodynamic favorability with increasing conductivity. Average ranged from 53 to 71 kJ mol C−1 (R2=0.78, p<0.00001), while no significant relationship between and specific conductivity was observed for the WSOC fraction in the deeper soils (averaging 51–54 kJ mol C−1) for the floodplain and inland samples (Fig. 3). The upland site had significantly higher average (67–70 kJ mol C−1) than the soils near the lowland. The (CHCl3) at both depths (Table S4) was higher than the water fractions (ranging between 96 and 105 kJ mol C−1) but did not show significant relationship with respect to specific conductivity.
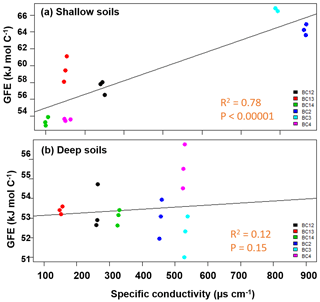
Figure 3Average Gibbs free energy (GFE) of samples in the water fraction of shallow soils impacted by tidal inundation increased with increasing specific conductivity (a), while no change was observed in the deeper soils (b). The salinity of the soil samples did not follow a clear spatial gradient.
Peak profiles for each solvent extraction showed distinct compound classes in the van Krevelen space, with peaks assigned to specific compound classes according to rules outlined in Kim et al. (2003) and modified by Bailey et al. (2017). The WSOC fraction was dominated by compounds classified as protein-, amino-sugar-, lignin-, condensed hydrocarbon-, carbohydrate-, and tannin-like compounds (Table 1), while the CHCl3 fraction had relative high abundances (75 % and higher) of lipid-like compounds (data not shown). A modest percentage of peaks (11 %–17 %) did not have classes assigned. Unique and common peaks extracted in the WSOC fraction in samples grouped according to their landscape position and depth (four sites in the floodplain (BC2, BC3, BC12, and BC13), two sites inland (BC4 and BC14), and one upland site (BC15)) are represented as H∕C to O∕C ratio of the compounds (p=0.05) in Fig. S1 in the Supplement.
Table 1Relative peak abundances (%) of compound classes in the water-extracted organic carbon fraction averaged across replicates per site. Samples are ordered according to their depth profile (shallow and deep) and their relative position in the landscape: floodplain (Fp), inland (I), and upland (U). Abbreviations: ConHC (condensed hydrocarbon), UnsatHC (unsaturated hydrocarbon), and Other (no classification assigned).
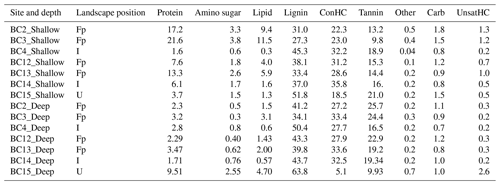
The shallow WSOC in the floodplain had greater relative abundance of unique lipid-like (28 %) and protein-like (25 %) compounds with relatively high H:C and low O:C ratios as compared to the upland site (BC15), which had 31 %, 30 %, and 19 % unique peaks, representing lignin-, tannin-, and carbohydrate-like compounds, respectively. About 23 % of peaks were common in both groups, including lignin- and condensed hydrocarbon-like compounds (Fig. S1a). Between the floodplain and the inland samples, high H:C and low O:C ratios representing 47 % lipid-, 38 % protein-, and 22 % amino-sugar-like peaks were uniquely present in the floodplain samples (Fig. S1b). The inland shallow soils had 19 % unique higher H:C peaks, representing condensed hydrocarbon-like compounds, compared to 1.2 % in the upland soil, though most of the compound classes were observed at both locations (Fig. S1c). Linear regression with specific conductivity profiles showed significant positive correlation with amino-sugar-, protein-, lipid-, and unsaturated hydrocarbon-like compounds, while condensed hydrocarbon-like compounds were significantly negatively correlated (Table S5).
For the deep soils, the upland site had 32 % unique peaks with relatively high H:C ratios and low O:C ratios commonly associated with unsaturated hydrocarbon-like compounds, as compared to the 0.7 % in the floodplain, which had higher prevalence of unique peaks, representing condensed hydrocarbon-like (36 %) and tannin-like (35 %) compounds (Table 1; Fig. S1d). The floodplain vs. inland samples had 3 times as many unique peaks, with high H:C and low O:C ratios representing lipid-like compounds in the floodplain samples. Comparisons between inland and upland endmember samples revealed 43 % and 37 % unique peaks, representing low H:C and high O:C ratio hydrocarbon- and tannin-like compounds, respectively, in inland samples, while 32 %, 14 %, 9 %, and 12 % of unique peaks were matched to unsaturated hydrocarbon-, lipid-, protein-, and amino-sugar-like compounds, respectively, in the latter (Table 1; Fig. S1e, f). No significant relationship between compound-class abundances and specific conductivity was observed (Table S5). For the CHCl3 fraction, peaks of lipid-like and unsaturated hydrocarbon-like compounds were observed to be common in all samples (data not shown), and regressions against specific conductivity were not significant for the compound classes.
Compositional differences of the organic compounds showed variable heteroatom abundances, with cumulative heteroatom abundance decreasing with increasing salinity (R2=0.43, p=0.009) for the shallow fraction of the WSOC. For the WSOC fraction, heteroatom abundance of CHOP (R2=0.61) and CHNOP (R2=0.50) containing compounds was consistent with our hypothesis and significantly (p<0.01) increased, while CHNOS (R2=0.66), and CHNOSP (R2=0.62) abundances were inconsistent with our hypothesis and significantly decreased with increasing specific conductivity. The elemental composition of the WSOC compounds for deep soils did not show any significant trend with respect to conductivity. For the CHCl3 fraction, relative abundance of CHNOP in the shallow soils significantly decreased with specific conductivity (R2=0.57, p<0.01).
3.3 Transformation profiles
In contrast to our expectations, the number of transformations decreased with increasing salinity in the water fraction of shallow soils (R2=0.60, p<0.01; Fig. 4a; Table S3). We also evaluated N-containing transformations and the abundance of N-containing compounds in the system. Total N-containing transformations also decreased significantly with conductivity, but the correlation was not as strong (R2=0.40, p<0.01). Total N-containing compounds (Fig. 4b; Table S3) as well as their relative abundance decreased significantly (R2=0.74, p<0.01), with increasing conductivity in the shallow soils for the water fraction.
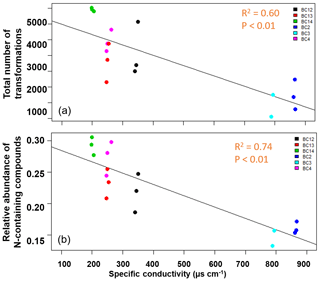
Figure 4The total number of inferred transformations (a) and total abundance of N-containing compounds (b) in the water fraction of shallow soils impacted by tidal inundation show significant negative correlations with increasing specific conductivity. No significant relationships were observed for water fraction of deeper soils or for the CHCl3 fraction in shallow or deeper soils. The salinity of the soil samples did not follow a clear spatial gradient.
3.4 Ecological processes impacting community composition
Null modeling revealed that microbial community assembly processes were influenced by variable selection (βNTI>2), homogenous selection (), dispersal limitation ( and RCbray>0.95), homogenizing dispersal ( and ), and undominated processes ( and ; Fig. 5). Dispersal limitation had the greatest influence, being responsible for 54 % of the variation in community composition. The lowest signal was for homogenizing dispersal (1 %), and the signal for homogenous selection (23 %) was higher than variable selection (9 %). Together, deterministic processes (variable selection plus homogeneous selection) were responsible for 32 % of the variation in community composition, with 55 % contributed by stochastic processes (dispersal limitation plus homogenizing dispersal). Variation not accounted for by dispersal or selection (i.e., influenced by a mixture of processes) accounted for the remaining signal (23 %). Consistent with influences from both stochastic and deterministic processes, βNTI relationships with environmental variables were significant (p<0.05 by the Mantel test) but relatively weak (r=0.46 for pH and r=0.31 for specific conductivity; Fig. S2).
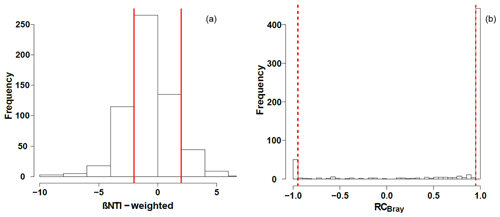
Figure 5Histograms representing the observed distribution of comparisons based on (a) β-nearest taxon index (βNTI) and (b) Raup–Crick metric (RCBray). Red lines represent the significance thresholds, whereby values outside of their bounds are significantly different from the null distribution.
To evaluate associations between microbial community assembly processes and chemistry, process estimates were regressed against features of the organic C profiles. Deterministic processes decreased (Fig. S3a), while community assembly processes influenced by non-deterministic processes increased with increasing number of transformations of organic compounds (Fig. S3b), although no strong relationships were observed between assembly processes and transformations (p=0.027 and R2=0.11 for deterministic or non-deterministic processes, p=0.475 and R2=0.015 for variable selection, p=0.054 and R2=0.10 for homogenous selection, p=0.514 and R2=0.013 for dispersal limitation, and p=0.627 and R2=0.007 for homogenizing dispersal). No significant relationships were observed between assembly processes and the number of N-containing transformations. Sørensen dissimilarity values based on the detected FTICR peaks for the water fraction were correlated with distance matrices of measured environmental variables and estimates of community assembly processes. Weak positive correlations were observed with NH4-N (r=0.28), pH (r=0.27), specific conductivity (r=0.41), NO3-N, silt, and clay (r=0.30), while for the CHCl3 fraction, weak positive correlations were observed with specific conductivity and NO3-N (r=0.26; Fig. S4). A Mantel test of FTICR Sørensen dissimilarity vs. βNTI values yielded a significant relationship (r=0.213, p=0.003) for the water fraction but not for the CHCl3 fraction (r=0.076, p=0.152). The stepwise model building yielded a combination of five variables that were weakly associated with the composition of water fraction peaks (p=0.026, adjusted R2=0.217), including sand, dispersal limitation, NH4-N concentration, specific conductivity, and location. The model explaining variation in the composition of the CHCl3 fraction peaks was non-significant (p=0.1, adjusted R2=0.05).
Sea level rise is increasing the inland extent of tides and exacerbating storm surge, resulting in greater salinity intrusion and altered ecosystem behavior across coastal TAIs (Conrads and Darby, 2017; Ensign and Noe, 2018; Langston et al., 2017; McCarthy et al., 2018; Neubauer et al., 2013). Site-driven variations in the responses of bulk soil biogeochemical processes (i.e., gas flux and DOC release) to elevated salinity suggest potentially important influences of underlying features such as C chemistry and microbial communities. To provide a foundation for understanding the role of C chemistry and microbial communities on biogeochemical cycling in coastal soils, we evaluated associations among a landscape-scale soil salinity gradient, molecular-level soil-carbon chemistry, and microbial community assembly processes in order to ultimately inform future improvements for predictive models. In soils associated with a coastal first-order drainage basin, we observed salinity-associated gradients in soil organic carbon fractions that were not associated with microbial community assembly processes. Our results are consistent with C chemistry being driven by a combination of spatially structured inputs driven by landscape structure (i.e., terrestrial inputs further inland and marine inputs further downstream) and salinity-associated metabolic responses of microbial communities that are independent of microbial community composition. An important caveat is that we did not measure microbial metabolism but instead infer an influence of microbial metabolism due to microbial composition being independent of C chemistry. To more directly evaluate these inferences, additional work is needed that focuses on quantifying inputs (e.g., via stable isotopes) and measuring microbial metabolism (e.g., via metatranscriptomics). Future work should also use tools like nuclear magnetic resonance and gas chromatography–mass spectrometry to evaluate low-molecular-weight OC (like those contributed by root exudates) that varies with salinity.
4.1 Molecular characterization reveals chemical gradients not seen in the bulk C pool
The systematic shifts observed in the molecular signatures compared to non-significant changes in bulk C chemistry show that molecular-level investigations are particularly relevant to process-based resolution of C biogeochemistry. The absence of bulk C signals mimicking molecular C signals parallel studies indicating rapid change in molecular constituents of the soil-C pool with no change in gross C content (Graham et al., 2018; Reynolds et al., 2018). A faster turnover time of C has been observed in microbial biomass as compared to bulk soil organic matter (Kramer and Gleixner, 2008), which is likely to impact microbe-mediated biochemical C transformations and lead to chemically complex heterogeneous C signatures likely to be missed in bulk analysis (Tfaily et al., 2015). The systematic shifts in chemical characteristics of soil-carbon fractions exhibited by samples at the shallow depth suggest that organic C compound pools in shallower soil depths are sensitive to salinity gradients, while deeper depth signatures do not vary systematically across the landscape. The landscape gradient observed in the shallow soils is likely influenced by a combination of reduced litterfall due to trees suffering under recent increases in salinity, changing understory vegetation, and algae-rich particulate OM deposition during inundation events that presumably initiated after the recent culvert removal (Wang et al., 2019). In contrast, the deeper soil depths were more similar to older organo-mineral complexed C in terrestrial soils across various ecosystems and land uses (Conant et al., 2011; Dungait et al., 2012; Jobbágy and Jackson, 2000; Kramer and Gleixner, 2006, 2008). The lack of any systematic gradients in the mineral-associated soil C provides further evidence in support of these interpretations, in addition to previous studies showing mineral-associated soil C to be less responsive to environmental forcings, relative to water-soluble C (Reynolds et al., 2018).
4.2 Decreases in organic C thermodynamic favorability may restrict microbial activity
Consistent with our first hypothesis, systematic changes in chemical characteristics of soil-carbon fractions were observed with thermodynamically less-favorable C present at high salinity in shallow soils. This gradient was expected to emerge from increased microbial activity at higher salinity, leaving behind less-favorable organic C. However, decreases in the number of inferred biochemical transformations and heteroatom abundances with increasing salinity suggests that microbial activity decreased with increasing salinity imply (but do not quantify) lower microbial activity at higher salinity. While difficult to infer direction of causality, these patterns suggest that less-favorable C at higher salinities may constrain microbial activity, leading to fewer biochemical transformations of the organic C. Thermodynamic limitation of organic C transformation is likely due to anaerobic conditions (LaRowe and Van Cappellen, 2011), which are indicated by high-moisture content of soils, high NH4-N, and low NO3-N. Anaerobic conditions restrict oxidation of C compounds based on thermodynamic properties (i.e., NOSC and ; Boye et al., 2017), and our data suggest that this has the potential to lead to lower microbial activity in conditions with less-favorable organic C.
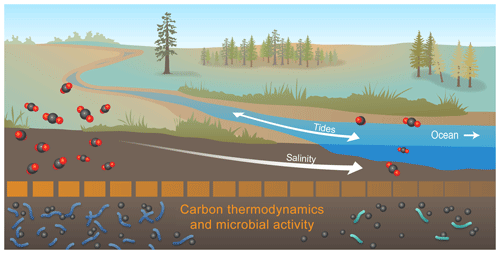
Figure 6Conceptual model summarizing the key outcome of our study: microbial activity at high salinity may be depressed by thermodynamically less-favorable C. Collectively, our data revealed that organic C thermodynamic favorability, heteroatom content, and number of biochemical transformations all decreased with increasing salinity. This suggests that microbial activity was lower at higher salinity, and we hypothesize that this was due to lower thermodynamic favorability of organic C. To evaluate generality, the salinity-associated gradients shown here need to be evaluated across coastal watersheds and mechanistically understood, as they have implications for contemporary and future C cycling in coastal watersheds experiencing hydrologic disturbances (e.g., sea level rise and storm surge).
4.3 Compound-class landscape gradients suggest influences of spatially structured inputs
Similar to patterns in C thermodynamic favorability, C compound classes showed significant heterogeneity in shallow soils but had conserved characteristics in deeper soils. The lipid-like peaks observed in the shallow floodplain samples suggest marine-associated algae-derived lipid organic matter similar to results observed by Ward et al. (2019b) in a coastal wetland setting. In contrast, lignin-like signatures in the upland site suggest terrestrially derived OM, as has been observed in other environments where terrestrially derived organic molecules have a high abundance of vascular-plant-derived material such as lignin (Hedges and Oades, 1997; Ward et al., 2013). These characteristics also align with reports of saturated soil environments (e.g., floodplains), exhibiting greater abundance of less-oxygenated organic matter than aerobic environments (e.g., upland soils), as reported by Tfaily et al. (2014) in organic matter transformation of a peat column. Our observed landscape gradients in compound-class composition indicate spatially structured inputs of organic C such as particulate OM deposition (Langley et al., 2007). Combining this outcome with gradients observed in the total number of biochemical transformations and the contribution of heteroatoms suggests that sources of C (marine vs. terrestrial) and in situ processing combine to influence landscape-scale gradients of molecular-level organic C chemistry.
4.4 Ecological assembly processes are weakly associated with organic C
Our results show that microbial community assembly is driven by a combination of dispersal limitation (a stochastic process) and deterministic selection most likely associated with pH, as is often observed in soils (Fierer, 2017; Fierer and Jackson, 2006; Garbeva et al., 2004). In contrast, variation in organic C character was associated primarily with specific conductivity. This suggests that the composition of microbial communities is not mechanistically related to C chemistry. Consistent with this inference, we found a very weak association between βNTI and organic C characteristics. Furthermore, and contrary to our hypothesis, we observed a weak negative association between the influence of deterministic processes and the number of organic C transformations.
Relatively fast changes of organic C chemistry compared to relatively slow changes in microbial composition may underlie the lack of association between assembly processes and C chemistry (Bramucci et al., 2013). Supporting this interpretation, a recent study evaluating microbial community composition and C biogeochemistry of soils in a mesohaline marsh following saltwater intrusion reported immediate changes in C mineralization rates with delayed shifts in microbial community composition (Dang et al., 2019). Similarly, a 17-year dryland soil transplant experiment showed large shifts in microbial activity with no change in community composition (Bond-Lamberty et al., 2016). Furthermore, studies across diverse systems show disconnect in function and composition. For example, C chemistry and not microbial community structure or gene expression was found to significantly influence freshwater hyporheic-zone organic matter processing (Graham et al., 2018). Environmental conditions influenced the distribution of functional groups, but not taxonomic composition of marine bacterial and archaeal communities (Lima-Mendez et al., 2015; Louca et al., 2016), and dynamic community shifts did not impact functional stability of a methanogenic reactor (Fernández et al., 1999). Combining our study with these previous investigations provides evidence that is consistent with (but does not prove) that soil microbial community composition can be independent of C chemistry, though this certainly varies across systems (e.g., Stegen et al., 2018b).
In our system, the lack of an association between microbial composition and organic C chemistry is also likely due to a strong influence of stochastic community assembly. Our null modeling indicated that dispersal limitation was responsible for 54 % of variation in community composition. Dispersal limitation influences composition by restricting the movement of organisms through space. Restricted movement enhances the influences of stochastic ecological drift, which arises through birth and death events that are randomly distributed across taxa (Green et al., 2004, 2008; Hubbell, 2001; Martiny et al., 2006; McClain et al., 2012; Stegen et al., 2015). Because ecological drift (enabled by dispersal limitation) can lead to the random loss of taxa within local communities, it can result in different communities containing different, but functionally redundant, taxa (Loreau, 2004). Moreover, one can argue, as per Louca et al. (2018), that in an open system with regular exposure to external inputs (e.g., via tides), functional redundancy is expected to occur and lead to a decoupling of microbial structure and function (Burke et al., 2011; Liebold and Chase, 2017; Nemergut et al., 2013).
Our results have revealed landscape-scale gradients in soil-C chemistry in a coastal forested floodplain, and they also show that such gradients are different across soil depths and OC fractions – occurring only in the shallow, water-soluble C pool. In addition, we found little evidence of an association between C chemistry and microbial community assembly processes, likely due to a dominant influence of stochastic community assembly (as indicated by a strong influence of dispersal limitation). We propose that the disconnect between C chemistry and microbial communities is enhanced by differences in the timescales for which C chemistry and microbial community composition shift.
Our findings suggest that cross-system heterogeneity observed in coastal soil biogeochemical responses to salinity is likely associated with molecular-level C chemistry and microbial physiological responses that are contingent on historical conditions (Fig. 6; Goldman et al., 2017; Hawkes and Keitt, 2015; Hawkes et al., 2017; Stegen et al., 2018a). We further suggest that microbial community composition may not strongly influence biogeochemical function in coastal soils. Processes associated with molecular-level C chemistry dynamics are therefore likely to be a critical component of ecosystem responses to changing salinity dynamics in coastal TAIs. A full elucidation of these processes will lay a foundation for the development of mechanistic models of coastal TAI biogeochemical dynamics, providing an opportunity for better representation of these ecosystems in local, regional, and Earth system models.
Raw sequence data have been uploaded to the National Center for Biotechnology Information's (NCBI's) Sequence Read Archive (SRA) under BioProject PRJNA541992. All other data are available at DataHub (a PNNL data repository), with digital objective identifiers at https://doi.org/10.25584/data.2019-08.931/1558461, https://doi.org/10.25584/data.2019-08.928/1558463, and https://doi.org/10.25584/data.2019-08.929/1558462 (Sengupta and Stegen, 2019). Original codes for community assembly metric calculation are available at the Stegen_etal_ISME 2013 GitHub repository at https://github.com/stegen/Stegen_etal_ISME_2013 (last access: 7 October 2019).
The supplement related to this article is available online at: https://doi.org/10.5194/bg-16-3911-2019-supplement.
AS designed the study, performed the experiments, conducted data analyses and interpretation, and wrote the original draft. JI and CG collected the samples and created site maps. MMT, RKC, and JT provided input on FTICR methodology, conducted the FTICR-MS instrument run, and handled quality filtering and pre-processing of FTICR scans. VLB and NDW contributed to funding acquisition, site selection, study design conceptualization, interpretation of results, and editing. JCS contributed to funding acquisition, study design conceptualization, interpretation of results, reviewing, and editing. All authors provided feedback on the paper.
The authors declare that they have no conflict of interest.
This work is part of the PREMIS Initiative at the Pacific Northwest National Laboratory (PNNL). It was funded and conducted under the Laboratory Directed Research and Development Program at PNNL, a multi-program national laboratory operated by Battelle for the US Department of Energy under Contract DE-AC05-76RL01830. A portion of the research was performed using EMSL (grid.436923.9), a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research. We would like to acknowledge Yuliya Farris and Sarah Fansler for DNA extraction and sequencing, respectively, Colin Brislawn for processing amplicon-sequence data, and the Central Analytical Laboratory at Oregon State University for conducting soil chemical analysis.
This paper was edited by Ji-Hyung Park and reviewed by two anonymous referees.
Ardón, M., Helton, A. M., and Bernhardt, E. S.: Drought and saltwater incursion synergistically reduce dissolved organic carbon export from coastal freshwater wetlands, Biogeochemistry, 127, 411–426, https://doi.org/10.1007/s10533-016-0189-5, 2016.
Ardón, M., Helton, A. M., and Bernhardt, E. S.: Salinity effects on greenhouse gas emissions from wetland soils are contingent upon hydrologic setting: a microcosm experiment, Biogeochemistry, 140, 217–232, https://doi.org/10.1007/s10533-018-0486-2, 2018.
Aronesty, E.: Comparison of sequencing utility programs, Open Bioinformatics Journal, 7, 1–8, 2013.
Bailey, V. L., Smith, A. P., Tfaily, M., Fansler, S. J., and Bond-Lamberty, B.: Differences in soluble organic carbon chemistry in pore waters sampled from different pore size domains, Soil Biol. Biochem., 107, 133–143, https://doi.org/10.1016/J.SOILBIO.2016.11.025, 2017.
Barry, S. C., Bianchi, T. S., Shields, M. R., Hutchings, J. A., Jacoby, C. A., and Frazer, T. K.: Characterizing blue carbon stocks in Thalassia testudinum meadows subjected to different phosphorus supplies: A lignin biomarker approach, Limnol. Oceanogr., 63, 2630–2646, https://doi.org/10.1002/lno.10965, 2018.
Bond-Lamberty, B., Bolton, H., Fansler, S., Heredia-Langner, A., Liu, C., McCue, L. A., Smith, J., and Bailey, V.: Soil Respiration and Bacterial Structure and Function after 17 Years of a Reciprocal Soil Transplant Experiment, edited by: Treseder, K., PLoS One, 11, e0150599, https://doi.org/10.1371/journal.pone.0150599, 2016.
Bottos, E. M., Kennedy, D. W., Romero, E. B., Fansler, S. J., Brown, J. M., Bramer, L. M., Chu, R. K., Tfaily, M. M., Jansson, J. K., and Stegen, J. C.: Dispersal limitation and thermodynamic constraints govern spatial structure of permafrost microbial communities, FEMS Microbiol. Ecol., 94, https://doi.org/10.1093/femsec/fiy110, 2018.
Boye, K., Noël, V., Tfaily, M. M., Bone, S. E., Williams, K. H., Bargar, J. R., and Fendorf, S.: Thermodynamically controlled preservation of organic carbon in floodplains, Nat. Geosci., 10, 415–419, https://doi.org/10.1038/NGEO2940, 2017.
Bramucci, A., Han, S., Beckers, J., Haas, C., Lanoil, B., Bramucci, A., Han, S., Beckers, J., Haas, C., and Lanoil, B.: Composition, Diversity, and Stability of Microbial Assemblages in Seasonal Lake Ice, Miquelon Lake, Central Alberta, Biology (Basel), 2, 514–532, https://doi.org/10.3390/biology2020514, 2013.
Breitling, R., Ritchie, S., Goodenowe, D., Stewart, M. L., and Barrett, M. P.: Ab initio prediction of metabolic networks using Fourier transform mass spectrometry data, Metabolomics, 2, 155–164, https://doi.org/10.1007/s11306-006-0029-z, 2006.
Brown, J., Zavoshy, N., Brislawn, C. J., and McCue, L. A.: Hundo: a Snakemake workflow for microbial community sequence data, PeerJ Preprints, 6, e27272v1, https://doi.org/10.7287/peerj.preprints.27272v1, 2018.
Burke, C., Steinberg, P., Rusch, D., Kjelleberg, S., and Thomas, T.: Bacterial community assembly based on functional genes rather than species, P. Natl. Acad. Sci. USA, 108, 14288–14293, https://doi.org/10.1073/pnas.1101591108, 2011.
Bushnell, B.: BBMap, available at: https://sourceforge.net/projects/bbmap/ (last access: 7 October 2019), 2018.
Caruso, T., Chan, Y., Lacap, D. C., Lau, M. C. Y., McKay, C. P., and Pointing, S. B.: Stochastic and deterministic processes interact in the assembly of desert microbial communities on a global scale, ISME J., 5, 1406–13, https://doi.org/10.1038/ismej.2011.21, 2011.
Chambers, L. G., Reddy, K. R., and Osborne, T. Z.: Short-Term Response of Carbon Cycling to Salinity Pulses in a Freshwater Wetland, Soil Sci. Soc. Am. J., 75, 2000, https://doi.org/10.2136/sssaj2011.0026, 2011.
Chambers, L. G., Osborne, T. Z., and Reddy, K. R.: Effect of salinity-altering pulsing events on soil organic carbon loss along an intertidal wetland gradient: a laboratory experiment, Biogeochemistry, 115, 363–383, https://doi.org/10.1007/s10533-013-9841-5, 2013.
Chambers, L. G., Davis, S. E., Troxler, T., Boyer, J. N., Downey-Wall, A., and Scinto, L. J.: Biogeochemical effects of simulated sea level rise on carbon loss in an Everglades mangrove peat soil, Hydrobiologia, 726, 195–211, https://doi.org/10.1007/s10750-013-1764-6, 2014.
Conant, R. T., Ogle, S. M., Paul, E. A., and Paustian, K.: Measuring and monitoring soil organic carbon stocks in agricultural lands for climate mitigation, Front. Ecol. Environ., 9, 169–173, https://doi.org/10.1890/090153, 2011.
Conrads, P. A. and Darby, L. S.: Development of a coastal drought index using salinity data, B. Am. Meteorol. Soc., 98, 753–766, https://doi.org/10.1175/BAMS-D-15-00171.1, 2017.
Dang, C., Morrissey, E. M., Neubauer, S. C., and Franklin, R. B.: Novel microbial community composition and carbon biogeochemistry emerge over time following saltwater intrusion in wetlands, Glob. Change Biol., 25, 549–561, https://doi.org/10.1111/gcb.14486, 2019.
Dini-Andreote, F., Stegen, J. C., van Elsas, J. D., and Salles, J. F.: Disentangling mechanisms that mediate the balance between stochastic and deterministic processes in microbial succession, P. Natl. Acad. Sci. USA, 112, E1326–1332, https://doi.org/10.1073/pnas.1414261112, 2015.
Dittmar, T., Koch, B., Hertkorn, N., and Kattner, G.: A simple and efficient method for the solid-phase extraction of dissolved organic matter (SPE-DOM) from seawater, Limnol. Oceanogr.-Meth., 6, 230–235, 2008.
Dungait, J. A. J., Hopkins, D. W., Gregory, A. S., and Whitmore, A. P.: Soil organic matter turnover is governed by accessibility not recalcitrance, Glob. Change Biol., 18, 1781–1796, https://doi.org/10.1111/j.1365-2486.2012.02665.x, 2012.
Edgar, R. C.: Search and clustering orders of magnitude faster than BLAST, Bioinformatics, 26, 2460–2461, 2010.
Ensign, S. H. and Noe, G. B.: Tidal extension and sea-level rise: recommendations for a research agenda, Front. Ecol. Environ., 16, 37–43, https://doi.org/10.1002/fee.1745, 2018.
ESRI: ArcGIS Desktop: Release 10.5 Redlands, Environmental Systems Research Institute, CA, 2017.
Fernández, A., Huang, S., Seston, S., Xing, J., Hickey, R., Criddle, C., and Tiedje, J.: How stable is stable? Function versus community composition, Appl. Environ. Microb., 65, 3697–704, 1999.
Fierer, N.: Embracing the unknown: disentangling the complexities of the soil microbiome, Nat. Publ. Gr., 15, 579–590, https://doi.org/10.1038/nrmicro.2017.87, 2017.
Fierer, N. and Jackson, R. B.: The diversity and biogeography of soil bacterial communities, P. Natl. Acad. Sci. USA, 103, 626–631, https://doi.org/10.1073/pnas.0507535103, 2006.
Garbeva, P., van Veen, J. A., and van Elsas, J. D.: Microbial diversity in soil: selection microbial populations by plant and soil type and implications for disease suppressiveness, Annu. Rev. Phytopathol., 42, 243–270, https://doi.org/10.1146/annurev.phyto.42.012604.135455, 2004.
Goldman, A. E., Graham, E. B., Crump, A. R., Kennedy, D. W., Romero, E. B., Anderson, C. G., Dana, K. L., Resch, C. T., Fredrickson, J. K., and Stegen, J. C.: Biogeochemical cycling at the aquatic–terrestrial interface is linked to parafluvial hyporheic zone inundation history, Biogeosciences, 14, 4229–4241, https://doi.org/10.5194/bg-14-4229-2017, 2017.
Gouffi, K., Pica, N., Pichereau, V., and Blanco, C.: Disaccharides as a new class of nonaccumulated osmoprotectants for Sinorhizobium meliloti, Appl. Environ. Microb., 65, 1491–1500, https://doi.org/10.1186/1746-1448-1-5, 1999.
Graham, E. B. and Stegen, J. C.: Stochastic microbial community assembly decreases biogeochemical function, bioRxiv, 183897, https://doi.org/10.1101/183897, 2017.
Graham, E. B., Knelman, J. E., Schindlbacher, A., Siciliano, S., Breulmann, M., Yannarell, A., Beman, J. M., Abell, G., Philippot, L., Prosser, J., Foulquier, A., Yuste, J. C., Glanville, H. C., Jones, D. L., Angel, R., Salminen, J., Newton, R. J., Bürgmann, H., Ingram, L. J., Hamer, U., Siljanen, H. M. P., Peltoniemi, K., Potthast, K., Bañeras, L., Hartmann, M., Banerjee, S., Yu, R.-Q., Nogaro, G., Richter, A., Koranda, M., Castle, S. C., Goberna, M., Song, B., Chatterjee, A., Nunes, O. C., Lopes, A. R., Cao, Y., Kaisermann, A., Hallin, S., Strickland, M. S., Garcia-Pausas, J., Barba, J., Kang, H., Isobe, K., Papaspyrou, S., Pastorelli, R., Lagomarsino, A., Lindström, E. S., Basiliko, N. and Nemergut, D. R.: Microbes as Engines of Ecosystem Function: When Does Community Structure Enhance Predictions of Ecosystem Processes?, Front. Microbiol., 7, 214, https://doi.org/10.3389/fmicb.2016.00214, 2016.
Graham, E. B., Tfaily, M. M., Crump, A. R., Goldman, A. E., Bramer, L. M., Arntzen, E., Romero, E., Resch, C. T., Kennedy, D. W., and Stegen, J. C.: Carbon Inputs From Riparian Vegetation Limit Oxidation of Physically Bound Organic Carbon Via Biochemical and Thermodynamic Processes, J. Geophys. Res.-Biogeo, 122, 3188–3205, https://doi.org/10.1002/2017JG003967, 2017a.
Graham, E. B., Crump, A. R., Resch, C. T., Fansler, S., Arntzen, E., Kennedy, D. W., Fredrickson, J. K., and Stegen, J. C.: Deterministic influences exceed dispersal effects on hydrologically-connected microbiomes, Environ. Microbiol., 19, 1552–1567, https://doi.org/10.1111/1462-2920.13720, 2017b.
Graham, E. B., Crump, A. R., Kennedy, D. W., Arntzen, E., Fansler, S., Purvine, S. O., Nicora, C. D., Nelson, W., Tfaily, M. M., and Stegen, J. C.: Multi'omics comparison reveals metabolome biochemistry, not microbiome composition or gene expression, corresponds to elevated biogeochemical function in the hyporheic zone, Sci. Total Environ., 642, 742–753, https://doi.org/10.1016/J.SCITOTENV.2018.05.256, 2018.
Green, J. L., Holmes, A. J., Westoby, M., Oliver, I., Briscoe, D., Dangerfield, M., Gillings, M., and Beattie, A. J.: Spatial scaling of microbial eukaryote diversity, Nature, 432, 747–750, https://doi.org/10.1038/nature03034, 2004.
Green, J. L., Bohannan, B. J. M., and Whitaker, R. J.: Microbial Biogeography: From Taxonomy to Traits, Science, 320, 1039–1043, https://doi.org/10.1126/SCIENCE.1153475, 2008.
Guillemette, R., Kaneko, R., Blanton, J., Tan, J., Witt, M., Hamilton, S., Allen, E. E., Medina, M., Hamasaki, K., Koch, B. P., and Azam, F.: Bacterioplankton drawdown of coral mass-spawned organic matter, ISME J., 12, 2238–2251, https://doi.org/10.1038/s41396-018-0197-7, 2018.
Hawkes, C. V. and Keitt, T. H.: Resilience vs. historical contingency in microbial responses to environmental change, edited by: Classen, A., Ecol. Lett., 18, 612–625, https://doi.org/10.1111/ele.12451, 2015.
Hawkes, C. V., Waring, B. G., Rocca, J. D., and Kivlin, S. N.: Historical climate controls soil respiration responses to current soil moisture, P. Natl. Acad. Sci. USA, 114, 6322–6327, https://doi.org/10.1073/pnas.1620811114, 2017.
Hedges, J. I. and Oades, J. M.: Comparative organic geochemistries of soils and marine sediments, Org. Geochem., 27, 319–361, 1997.
Hedges, J. I., Keil, R. G., and Benner, R.: What happens to terrestrial organic matter in the ocean?, Org. Geochem., 27, 195–212, https://doi.org/10.1016/S0146-6380(97)00066-1, 1997.
Herbert, E. R., Schubauer-Berigan, J., and Craft, C. B.: Differential effects of chronic and acute simulated seawater intrusion on tidal freshwater marsh carbon cycling, Biogeochemistry, 138, 137–154, https://doi.org/10.1007/s10533-018-0436-z, 2018.
Hinson, A. L., Feagin, R. A., Eriksson, M., Najjar, R. G., Herrmann, M., Bianchi, T. S., Kemp, M., Hutchings, J. A., Crooks, S., and Boutton, T.: The spatial distribution of soil organic carbon in tidal wetland soils of the continental United States, Glob. Change Biol., 23, 5468–5480, https://doi.org/10.1111/gcb.13811, 2017.
Hoitink, A. J. F. and Jay, D. A.: Tidal river dynamics: Implications for deltas, Rev. Geophys., 54, 240–272, https://doi.org/10.1002/2015RG000507, 2016.
Hoitink, A. J. F., Buschman, F. A., and Vermeulen, B.: Continuous measurements of discharge from a horizontal acoustic Doppler current profiler in a tidal river, Water Resour. Res, 45, 11406, https://doi.org/10.1029/2009WR007791, 2009.
Holmquist, J. R., Windham-Myers, L., Bliss, N., Crooks, S., Morris, J. T., Megonigal, J. P., Troxler, T., Weller, D., Callaway, J., Drexler, J., Ferner, M. C., Gonneea, M. E., Kroeger, K. D., Schile-Beers, L., Woo, I., Buffington, K., Breithaupt, J., Boyd, B. M., Brown, L. N., Dix, N., Hice, L., Horton, B. P., MacDonald, G. M., Moyer, R. P., Reay, W., Shaw, T., Smith, E., Smoak, J. M., Sommerfield, C., Thorne, K., Velinsky, D., Watson, E., Grimes, K. W., and Woodrey, M.: Accuracy and Precision of Tidal Wetland Soil Carbon Mapping in the Conterminous United States, Sci. Rep.-UK, 8, 9478, https://doi.org/10.1038/s41598-018-26948-7, 2018.
Hubbell, S. P.: The Unified Neutral Theory of Biodiversity and Biogeography, Princeton University Press, available at: http://www.jstor.org/stable/j.ctt7rj8w (last access: 7 October 2019), 2001.
Jobbágy, E. G. and Jackson, R. B.: The vertical distribution of soil organic carbon and its relation to climate and vegetation, Ecol. Appl., 10, 423–436, https://doi.org/10.1890/1051-0761(2000)010[0423:TVDOSO]2.0.CO;2, 2000.
Kim, S., Kramer, R. W., and Hatcher, P. G.: Graphical Method for Analysis of Ultrahigh-Resolution Broadband Mass Spectra of Natural Organic Matter, the Van Krevelen Diagram, Anal. Chem., 75, 5336–5344, https://doi.org/10.1021/ac034415p, 2003.
Koch, B. P. and Dittmar, T.: From mass to structure: an aromaticity index for high-resolution mass data of natural organic matter, Rapid Commun. Mass Sp., 20, 926–932, https://doi.org/10.1002/rcm.2386, 2006.
Koch, B. P. and Dittmar, T.: From mass to structure: an aromaticity index for high-resolution mass data of natural organic matter, Rapid Commun. Mass Sp., 30, 250–250, https://doi.org/10.1002/rcm.7433, 2016.
Koch, B. P., Kattner, G., Witt, M., and Passow, U.: Molecular insights into the microbial formation of marine dissolved organic matter: recalcitrant or labile?, Biogeosciences, 11, 4173–4190, https://doi.org/10.5194/bg-11-4173-2014, 2014.
Kramer, C. and Gleixner, G.: Variable use of plant- and soil-derived carbon by microorganisms in agricultural soils, Soil Biol. Biochem., 38, 3267–3278, https://doi.org/10.1016/J.SOILBIO.2006.04.006, 2006.
Kramer, C. and Gleixner, G.: Soil organic matter in soil depth profiles: Distinct carbon preferences of microbial groups during carbon transformation, Soil Biol. Biochem., 40, 425–433, https://doi.org/10.1016/J.SOILBIO.2007.09.016, 2008.
Krauss, K. W., Noe, G. B., Duberstein, J. A., Conner, W. H., Stagg, C. L., Cormier, N., Jones, M. C., Bernhardt, C. E., Graeme Lockaby, B., From, A. S., Doyle, T. W., Day, R. H., Ensign, S. H., Pierfelice, K. N., Hupp, C. R., Chow, A. T., and Whitbeck, J. L.: The Role of the Upper Tidal Estuary in Wetland Blue Carbon Storage and Flux, Global Biogeochem. Cy., 32, 817–839, https://doi.org/10.1029/2018GB005897, 2018.
Ksionzek, K. B., Lechtenfeld, O. J., McCallister, S. L., Schmitt-Kopplin, P., Geuer, J. K., Geibert, W., and Koch, B. P.: Dissolved organic sulfur in the ocean: Biogeochemistry of a petagram inventory, Science, 354, 456–459, https://doi.org/10.1126/science.aaf7796, 2016.
Kubartová, A., Ottosson, E., and Stenlid, J.: Linking fungal communities to wood density loss after 12 years of log decay, FEMS Microbiol. Ecol., 91, fiv032, https://doi.org/10.1093/femsec/fiv032, 2015.
Kujawinski, E. B. and Behn, M. D.: Automated analysis of electrospray ionization fourier transform ion cyclotron resonance mass spectra of natural organic matter, Anal. Chem., 78, 4363–73, https://doi.org/10.1021/ac0600306, 2006.
Langer, U. and Rinklebe, J.: Lipid biomarkers for assessment of microbial communities in floodplain soils of the Elbe River (Germany), Wetlands, 29, 353–362, https://doi.org/10.1672/08-114.1, 2009.
Langley, J. A., Sigrist, M. V., Duls, J., Cahoon, D. R., Lynch, J. C., and Megonigal, J. P.: Global Change and Marsh Elevation Dynamics: Experimenting Where Land Meets Sea and Biology Meets Geology, in: Proceedings of the Smithsonian Marine Sciences Symposium, 391–400, Smithsonian Institution Scholarly Press, Washington, DC, available at: https://repository.si.edu/bitstream/handle/10088/18988/serc_Langley_et_al._2009_Smithsonian_Marine_Sciences_.pdf?sequence=1&isAllowed=y (last access: 7 May 2019), 2007.
Langston, A. K., Kaplan, D. A., and Putz, F. E.: A casualty of climate change? Loss of freshwater forest islands on Florida's Gulf Coast, Glob. Change Biol., 23, 5383–5397, https://doi.org/10.1111/gcb.13805, 2017.
LaRowe, D. E. and Van Cappellen, P.: Degradation of natural organic matter: A thermodynamic analysis, Geochim. Cosmochim. Ac., 75, 2030–2042, https://doi.org/10.1016/J.GCA.2011.01.020, 2011.
Lechtenfeld, O. J., Hertkorn, N., Shen, Y., Witt, M., and Benner, R.: Marine sequestration of carbon in bacterial metabolites., Nat. Commun., 6, 6711, https://doi.org/10.1038/ncomms7711, 2015.
Liebold, M. A. and Chase, M. J.: Combining Taxonomic and Functional Patterns to Disentangle Metacommunity Assembly Processes, in: Metacommunity Ecology, Vol. 9, edited by: Liebold, M. A. and Chase, M. J., p. 504, Princeton University Press, Princeton, New Jersey, 2017.
Lima-Mendez, G., Faust, K., Henry, N., Decelle, J., Colin, S., Carcillo, F., Chaffron, S., Ignacio-Espinosa, J. C., Roux, S., Vincent, F., Bittner, L., Darzi, Y., Wang, J., Audic, S., Berline, L., Bontempi, G., Cabello, A. M., Coppola, L., Cornejo-Castillo, F. M., d'Ovidio, F., Meester, L. De, Ferrera, I., Garet-Delmas, M.-J., Guidi, L., Lara, E., Pesant, S., Royo-Llonch, M., Salazar, G., Sánchez, P., Sebastian, M., Souffreau, C., Dimier, C., Picheral, M., Searson, S., Kandels-Lewis, S., TARA Oceans coordinators, Gorsky, G., Not, F., Ogata, H., Speich, S., Stemmann, L., Weissenbach, J., Wincker, P., Acinas, S. G., Sunagawa, S., Bork, P., Sullivan, M. B., Karsenti, E., Bowler, C., Vargas, C. de, and Raes, J.: Determinants of community structure in the global plankton interactome, Science, 348, 1262073, https://doi.org/10.1126/SCIENCE.1262073, 2015.
Liu, X., Ruecker, A., Song, B., Xing, J., Conner, W. H., and Chow, A. T.: Effects of salinity and wet–dry treatments on C and N dynamics in coastal-forested wetland soils: Implications of sea level rise, Soil Biol. Biochem., 112, 56–67, https://doi.org/10.1016/J.SOILBIO.2017.04.002, 2017.
Loreau, M.: Does functional redundancy exist?, Oikos, 104, 606–611, https://doi.org/10.1111/j.0030-1299.2004.12685.x, 2004.
Louca, S., Parfrey, L. W., and Doebeli, M.: Decoupling function and taxonomy in the global ocean microbiome, Science, 353, 1272–1277, https://doi.org/10.1126/science.aaf4507, 2016.
Louca, S., Polz, M. F., Mazel, F., Albright, M. B. N., Huber, J. A., O'connor, M. I., Ackermann, M., Hahn, A. S., Srivastava, D. S., Crowe, S. A., Doebeli, M., and Parfrey, L. W.: Disentangling function from taxonomy in microbial systems Function and functional redundancy in microbial systems, Nature Ecology & Evolution, 2, 936–943, https://doi.org/10.1038/s41559-018-0519-1, 2018.
Martiny, J. B. H., Bohannan, B. J. M., Brown, J. H., Colwell, R. K., Fuhrman, J. A., Green, J. L., Horner-Devine, M. C., Kane, M., Krumins, J. A., Kuske, C. R., Morin, P. J., Naeem, S., Øvreås, L., Reysenbach, A.-L., Smith, V. H., and Staley, J. T.: Microbial biogeography: putting microorganisms on the map, Nat. Rev. Microbiol., 4, 102–112, https://doi.org/10.1038/nrmicro1341, 2006.
Marton, J. M., Herbert, E. R., and Craft, C. B.: Effects of Salinity on Denitrification and Greenhouse Gas Production from Laboratory-incubated Tidal Forest Soils, Wetlands, 32, 347–357, https://doi.org/10.1007/s13157-012-0270-3, 2012.
Mau, R. L., Liu, C. M., Aziz, M., Schwartz, E., Dijkstra, P., Marks, J. C., Price, L. B., Keim, P., and Hungate, B. A.: Linking soil bacterial biodiversity and soil carbon stability, ISME J., 9, 1477–1480, https://doi.org/10.1038/ismej.2014.205, 2015.
McCarthy, M., Dimmitt, B., Muller-Karger, F., McCarthy, M. J., Dimmitt, B., and Muller-Karger, F. E.: Rapid Coastal Forest Decline in Florida's Big Bend, Remote Sens., 10, 1721, https://doi.org/10.3390/rs10111721, 2018.
McClain, C. R., Stegen, J. C., and Hurlbert, A. H.: Dispersal, environmental niches and oceanic-scale turnover in deep-sea bivalves, P. Roy. Soc. B-Biol. Sci., 279, https://doi.org/10.1098/RSPB.2011.2166, 2012.
Medeiros, P. M., Seidel, M., Ward, N. D., Carpenter, E. J., Gomes, H. R., Niggemann, J., Krusche, A. V., Richey, J. E., Yager, P. L., and Dittmar, T.: Fate of the Amazon River dissolved organic matter in the tropical Atlantic Ocean, Global Biogeochem. Cy., 29, 677–690, https://doi.org/10.1002/2015GB005115, 2015.
Minor, E. C., Steinbring, C. J., Longnecker, K., and Kujawinski, E. B.: Characterization of dissolved organic matter in Lake Superior and its watershed using ultrahigh resolution mass spectrometry, Org. Geochem., 43, 1–11, https://doi.org/10.1016/J.ORGGEOCHEM.2011.11.007, 2012.
Nemergut, D. R., Schmidt, S. K., Fukami, T., O'Neill, S. P., Bilinski, T. M., Stanish, L. F., Knelman, J. E., Darcy, J. L., Lynch, R. C., Wickey, P., and Ferrenberg, S.: Patterns and processes of microbial community assembly, Microbiol. Mol. Biol. R., 77, 342–56, https://doi.org/10.1128/MMBR.00051-12, 2013.
Neubauer, S. C., Givler, K., Valentine, S., and Megonigal, J. P.: Seasonal patterns and plant-mediated controls of subsurface wetland biogeochemistry, Ecology, 86, 3334–3344, https://doi.org/10.1890/04-1951, 2005.
Neubauer, S. C., Franklin, R. B., and Berrier, D. J.: Saltwater intrusion into tidal freshwater marshes alters the biogeochemical processing of organic carbon, Biogeosciences, 10, 8171–8183, https://doi.org/10.5194/bg-10-8171-2013, 2013.
Nyman, J. A. and Delaune, R. D.: CO, emission and soil Eh responses to different hydrological conditions in fresh, brackish, and saline marsh soils, available at: https://aslopubs.onlinelibrary.wiley.com/doi/pdf/10.4319/lo.1991.36.7.1406 (last access: 31 October 2018), 1991.
R Development Core Team: The R Project for Statistical Computing, available at: https://www.r-project.org/, last access: 9 May 2019.
Reynolds, L. L., Lajtha, K., Bowden, R. D., Tfaily, M. M., Johnson, B. R., and Bridgham, S. D.: The Path From Litter to Soil: Insights Into Soil C Cycling From Long-Term Input Manipulation and High-Resolution Mass Spectrometry, J. Geophys. Res.-Biogeo., 123, 1486–1497, https://doi.org/10.1002/2017JG004076, 2018.
Rivas-Ubach, A., Liu, Y., Bianchi, T. S., Tolić, N., Jansson, C., and Paša-Tolić, L.: Moving beyond the van Krevelen Diagram: A New Stoichiometric Approach for Compound Classification in Organisms, Anal. Chem., 90, 6152–6160, https://doi.org/10.1021/acs.analchem.8b00529, 2018.
Rocca, J. D., Hall, E. K., Lennon, J. T., Evans, S. E., Waldrop, M. P., Cotner, J. B., Nemergut, D. R., Graham, E. B., and Wallenstein, M. D.: Relationships between protein-encoding gene abundance and corresponding process are commonly assumed yet rarely observed, ISME J., 9, 1693–1699, https://doi.org/10.1038/ismej.2014.252, 2015.
Sawakuchi, H. O., Neu, V., Ward, N. D., Barros, M. de L. C., Valerio, A. M., Gagne-Maynard, W., Cunha, A. C., Less, D. F. S., Diniz, J. E. M., Brito, D. C., Krusche, A. V., and Richey, J. E.: Carbon Dioxide Emissions along the Lower Amazon River, Front. Mar. Sci., 4, 76, https://doi.org/10.3389/fmars.2017.00076, 2017.
Schmidt, M. W. I., Torn, M. S., Abiven, S., Dittmar, T., Guggenberger, G., Janssens, I. A., Kleber, M., Kögel-Knabner, I., Lehmann, J., Manning, D. A. C., Nannipieri, P., Rasse, D. P., Weiner, S., and Trumbore, S. E.: Persistence of soil organic matter as an ecosystem property, Nature, 478, 49–56, https://doi.org/10.1038/nature10386, 2011.
Seidel, M., Dittmar, T., Ward, N. D., Krusche, A. V., Richey, J. E., Yager, P. L., and Medeiros, P. M.: Seasonal and spatial variability of dissolved organic matter composition in the lower Amazon River, Biogeochemistry, 131, 281–302, https://doi.org/10.1007/s10533-016-0279-4, 2016.
Sengupta, A. and Stegen, J. C.: PREMIS – Terrestrial-Aquatic Interface, DataHub, https://datahub.pnnl.gov/datahub/project/8 (last access: 8 October 2019), 2019.
Sengupta, A., Stegen, J. C., Meira Neto, A. A., Wang, Y., Neilson, J. W., Chorover, J., Troch, P. A., Maier, R. M., Chorover, J., Troch, P. A., and Maier, R. M.: Assessing Microbial Community Patterns During Incipient Soil Formation From Basalt, J. Geophys. Res.-Biogeo., 124, 941–958, https://doi.org/10.1029/2017JG004315, 2019.
Shen, Y., Chapelle, F. H., Strom, E. W., and Benner, R.: Origins and bioavailability of dissolved organic matter in groundwater, Biogeochemistry, 122, 61–78, https://doi.org/10.1007/s10533-014-0029-4, 2015.
Simon, C., Roth, V.-N., Dittmar, T., and Gleixner, G.: Molecular Signals of Heterogeneous Terrestrial Environments Identified in Dissolved Organic Matter: A Comparative Analysis of Orbitrap and Ion Cyclotron Resonance Mass Spectrometers, Front. Earth Sci., 6, 138, https://doi.org/10.3389/feart.2018.00138, 2018.
Sleator, R. D. and Hill, C.: Bacterial osmoadaptation: The role of osmolytes in bacterial stress and virulence, FEMS Microbiol. Rev., 26, 49–71, https://doi.org/10.1016/S0168-6445(01)00071-7, 2002.
Smith, C. J., DeLaune, R. D., and Patrick, W. H.: Carbon dioxide emission and carbon accumulation in coastal wetlands, Estuar. Coast. Shelf S., 17, 21–29, https://doi.org/10.1016/0272-7714(83)90042-2, 1983.
Stegen, J. C., Lin, X., Konopka, A. E., and Fredrickson, J. K.: Stochastic and deterministic assembly processes in subsurface microbial communities, ISME J., 6, 1653–1664, https://doi.org/10.1038/ismej.2012.22, 2012.
Stegen, J. C., Lin, X., Fredrickson, J. K., Chen, X., Kennedy, D. W., Murray, C. J., Rockhold, M. L., and Konopka, A.: Quantifying community assembly processes and identifying features that impose them, ISME J., 7, 2069–2079, https://doi.org/10.1038/ismej.2013.93, 2013.
Stegen, J. C., Lin, X., Fredrickson, J. K., and Konopka, A. E.: Estimating and mapping ecological processes influencing microbial community assembly, Front. Microbiol., 6, 370, https://doi.org/10.3389/fmicb.2015.00370, 2015.
Stegen, J. C., Fredrickson, J. K., Wilkins, M. J., Konopka, A. E., Nelson, W. C., Arntzen, E. V., Chrisler, W. B., Chu, R. K., Danczak, R. E., Fansler, S. J., Kennedy, D. W., Resch, C. T., and Tfaily, M.: Groundwater–surface water mixing shifts ecological assembly processes and stimulates organic carbon turnover, Nat. Commun., 7, 11237, https://doi.org/10.1038/ncomms11237, 2016.
Stegen, J. C., Bottos, E. M., and Jansson, J. K.: A unified conceptual framework for prediction and control of microbiomes, Curr. Opin. Microbiol., 44, 20–27, https://doi.org/10.1016/J.MIB.2018.06.002, 2018a.
Stegen, J. C., Johnson, T., Fredrickson, J. K., Wilkins, M. J., Konopka, A. E., Nelson, W. C., Arntzen, E. V., Chrisler, W. B., Chu, R. K., Fansler, S. J., Graham, E. B., Kennedy, D. W., Resch, C. T., Tfaily, M., and Zachara, J.: Influences of organic carbon speciation on hyporheic corridor biogeochemistry and microbial ecology, Nat. Commun., 9, 1–11, https://doi.org/10.1038/s41467-018-02922-9, 2018b.
Steinmuller, H. E. and Chambers, L. G.: Can Saltwater Intrusion Accelerate Nutrient Export from Freshwater Wetland Soils? An Experimental Approach, Soil Sci. Soc. Am. J., 82, 283, https://doi.org/10.2136/sssaj2017.05.0162, 2018.
Sumner, M. E.: Handbook of Soil Science, edited by: Sumner, M. E., CRC Press, New York, 1999.
Tank, S. E., Fellman, J. B., Hood, E., and Kritzberg, E. S.: Beyond respiration: Controls on lateral carbon fluxes across the terrestrial-aquatic interface, Limnol. Oceanogr. Lett., 3, 76–88, https://doi.org/10.1002/lol2.10065, 2018.
Tfaily, M. M., Hodgkins, S., Podgorski, D. C., Chanton, J. P., and Cooper, W. T.: Comparison of dialysis and solid-phase extraction for isolation and concentration of dissolved organic matter prior to Fourier transform ion cyclotron resonance mass spectrometry, Anal. Bioanal. Chem., 404, 447–457, https://doi.org/10.1007/s00216-012-6120-6, 2012.
Tfaily, M. M., Cooper, W. T., Kostka, J. E., Chanton, P. R., Schadt, C. W., Hanson, P. J., Iversen, C. M., and Chanton, J. P.: Organic matter transformation in the peat column at Marcell Experimental Forest: Humification and vertical stratification, J. Geophys. Res.-Biogeo., 119, 661–675, https://doi.org/10.1002/2013JG002492, 2014.
Tfaily, M. M., Chu, R. K., Tolić, N., Roscioli, K. M., Anderton, C. R., Paša-Tolić, L., Robinson, E. W., and Hess, N. J.: Advanced Solvent Based Methods for Molecular Characterization of Soil Organic Matter by High-Resolution Mass Spectrometry, Anal. Chem., 87, 5206–5215, https://doi.org/10.1021/acs.analchem.5b00116, 2015.
Tfaily, M. M., Chu, R. K., Toyoda, J., Tolić, N., Robinson, E. W., Paša-Tolić, L., and Hess, N. J.: Sequential extraction protocol for organic matter from soils and sediments using high resolution mass spectrometry, Anal. Chim. Acta, 972, 54–61, https://doi.org/10.1016/J.ACA.2017.03.031, 2017.
Tolić, N., Liu, Y., Liyu, A., Shen, Y., Tfaily, M. M., Kujawinski, E. B., Longnecker, K., Kuo, L.-J., Robinson, E. W., Paša-Tolić, L., and Hess, N. J.: Formularity: Software for Automated Formula Assignment of Natural and Other Organic Matter from Ultrahigh-Resolution Mass Spectra, Anal. Chem., 89, 12659–12665, https://doi.org/10.1021/acs.analchem.7b03318, 2017.
Trivedi, P., Delgado-Baquerizo, M., Trivedi, C., Hu, H., Anderson, I. C., Jeffries, T. C., Zhou, J., and Singh, B. K.: Microbial regulation of the soil carbon cycle: evidence from gene–enzyme relationships, ISME J., 10, 2593–2604, https://doi.org/10.1038/ismej.2016.65, 2016.
Tzortziou, M., Neale, P. J., Megonigal, P., Pow, C. L., and Butterworth, M.: Spatial gradients in dissolved carbon due to tidal marsh outwelling into a Chesapeake Bay estuary, Mar. Ecol.-Prog. Ser., 426, 41–56, https://doi.org/10.3354/meps09017, 2011.
U.S. DOE: Research Priorities to Incorporate Terrestrial-Aquatic Interfaces in Earth System Models: Workshop Report, DOE/SC-0187, 2017.
van der Wal, A., Ottosson, E., and de Boer, W.: Neglected role of fungal community composition in explaining variation in wood decay rates, Ecology, 96, 124–133, https://doi.org/10.1890/14-0242.1, 2015.
Vidon, P., Allan, C., Burns, D., Duval, T. P., Gurwick, N., Inamdar, S., Lowrance, R., Okay, J., Scott, D., and Sebestyen, S.: Hot Spots and Hot Moments in Riparian Zones: Potential for Improved Water Quality Management1, J. Am. Water Resour. As., 46, 278–298, https://doi.org/10.1111/j.1752-1688.2010.00420.x, 2010.
Wang, W., McDowell, N. G., Ward, N. D., Indivero, J., Gunn, C., and Bailey, V. L.: Constrained tree growth and gas‐exchange of seawater exposed forests in the Pacific Northwest, USA, J. Ecol., https://doi.org/10.1111/1365-2745.13225, online first, 2019.
Ward, C. P., Nalven, S. G., Crump, B. C., Kling, G. W., and Cory, R. M.: Photochemical alteration of organic carbon draining permafrost soils shifts microbial metabolic pathways and stimulates respiration, Nat. Commun., 8, 772, https://doi.org/10.1038/s41467-017-00759-2, 2017.
Ward, N. D., Keil, R. G., Medeiros, P. M., Brito, D. C., Cunha, A. C., Dittmar, T., Yager, P. L., Krusche, A. V., and Richey, J. E.: Degradation of terrestrially derived macromolecules in the Amazon River, Nat. Geosci., 6, 530–533, https://doi.org/10.1038/ngeo1817, 2013.
Ward, N. D., Bianchi, T. S., Medeiros, P. M., Seidel, M., Richey, J. E., Keil, R. G., and Sawakuchi, H. O.: Where Carbon Goes When Water Flows: Carbon Cycling across the Aquatic Continuum, Front. Mar. Sci., 4, 7, https://doi.org/10.3389/fmars.2017.00007, 2017.
Ward, N. D., Indivero, J., Gunn, C., Wang, W., Bailey, V., and McDowell, N. G.: Longitudinal gradients in tree stem greenhouse gas concentrations across six Pacific Northwest coastal forests, J. Geophys. Res.-Biogeo., 124, 1401–1412, https://doi.org/10.1029/2019JG005064, 2019a.
Ward, N. D., Morrison, E. S., Liu, Y., Rivas-Ubach, A., Osborne, T. Z., Ogram, A. V., and Bianchi, T. S.: Marine microbial community responses related to wetland carbon mobilization in the coastal zone, Limnol. Oceanogr. Lett., 4, 25–33, https://doi.org/10.1002/lol2.10101, 2019b.
Washington Department of Fish and Wildlife: Fish Passage & Diversion Screening Inventory, available at: https://geo.wa.gov/datasets/4477faa67e95467cb5d3db75f41e7291_0?geometry=-135.166%2C47.551%2C-107.414%2C52.487 , last access: 7 October 2019.
Weston, N. B., Dixon, R. E., and Joye, S. B.: Ramifications of increased salinity in tidal freshwater sediments: Geochemistry and microbial pathways of organic matter mineralization, J. Geophys. Res., 111, G01009, https://doi.org/10.1029/2005JG000071, 2006.
Weston, N. B., Vile, M. A., Neubauer, S. C., and Velinsky, D. J.: Accelerated microbial organic matter mineralization following salt-water intrusion into tidal freshwater marsh soils, Biogeochemistry, 102, 135–151, https://doi.org/10.1007/s10533-010-9427-4, 2011.
Weston, N. B., Neubauer, S. C., Velinsky, D. J., and Vile, M. A.: Net ecosystem carbon exchange and the greenhouse gas balance of tidal marshes along an estuarine salinity gradient, Biogeochemistry, 120, 163–189, https://doi.org/10.1007/s10533-014-9989-7, 2014.
Yang, J., Zhan, C., Li, Y., Zhou, D., Yu, Y., and Yu, J.: Effect of salinity on soil respiration in relation to dissolved organic carbon and microbial characteristics of a wetland in the Liaohe River estuary, Northeast China, Sci. Total Environ., 642, 946–953, https://doi.org/10.1016/J.SCITOTENV.2018.06.121, 2018.
Zark, M. and Dittmar, T.: Universal molecular structures in natural dissolved organic matter, Nat. Commun., 9, 3178, https://doi.org/10.1038/s41467-018-05665-9, 2018.





